PFM2PWM: which “nucleotide background frequency” to use
13th September 2006
As I previously mentioned, in converting PFM to PWM single variable – [prior] background nucleotide frequency – was ambiguous to me. From other articles I noticed that it is usually set to 0.25 (1/4 – because there 4 nucleotides, thus in “perfectly random” sequence they would appear in 25% of cases each). In that post, I also thought of using “real” background frequency of nucleotides, calculated from the sequence, to which the matrix is to be applied.
I wrote a program to search all the 1000-basepair upstream sequences from all human, rat and mouse genes, present in Ensembl database release 40 (assuming those 1kb upstreams to be “promoters” of genes). For each promoter, only the single best score was returned. Then I draw a graph of the distribution of the number of promoters (y-axis) depending on the best match scores (x-axis). I did the search twice – one with p(b) = 0.25, and one with p(b) set to calculated values of A/C/G/T content in each promoter.
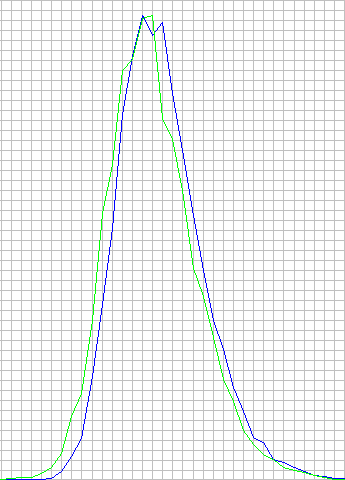
x-axis: similarity score, %, from 0 to 100
y-axis: number of promoters, which had best match with that similarity score (absolute number value)
1 box is 1% on x-axis. Blue color is for p(b) calculated from sequence, green color is for p(b) = 0.25.
It can be clearly seen, that both graphs are nearly the same, with the only noticeable difference in green graph being shifted leftwards a little bit (1-2%%). The same search was performed against a set of all exons for three above-mentioned organisms (exons also from Ensembl). The difference was nearly the same there as well – p(b) = 0.25 was slightly shifted to the left.
Actually, this behaviour is expected: by providing sequence-optimized p(b), we allow better similarity scores for matrix-sequence matching.
Anyway, providing “experimental evidence” is better than using “expected behaviour”. As an answer to the “which frequency to use” question I would recommend using sequence-optimized p(b). After I do usual statistical treatment of the data I got, I will publish the std-based recommendation. Stay tuned ![]()

October 18th, 2006 at 23:40
[...] PFM2PWM: which “nucleotide background frequency” to use [...]